Building blocks of heredity and behavior: Genes, DNA, RNA, proteins and co
Here we present the elements of genetic information transfer in a very condensed form. This article is based on various - much more detailed - articles from the German Wikipedia and The Sequence Onotolgy.
For the neurological basics (neurons, glial cells, synapses, neurotransmitters, hormones, receptors, action potential, blood-brain barrier see under Neurological basics
- 1. Gene
- 2. DNA (deoxyribonucleic acid)
- 3. RNA (ribonucleic acid)
- 4. Nucleotides
- 5. Proteins (proteins)
- 6. Amino acids (aminocarboxylic acids)
1. Gene
A gene is a section of DNA.
The genes in the DNA contain the information for the production of RNA (ribonucleic acids).
Protein-coding genes encode mRNA (messenger RNA). The mRNA contains the information for the construction of proteins.
The sequence of bases determines the sequence of amino acids in the protein. Three adjacent nucleotides form a codon, which can be used to uniquely determine a specific amino acid that is to be incorporated into a protein.
1.1. Gene expression
The expression of a gene describes its activity to form the gene product encoded by the gene (in particular proteins or RNA molecules).
The expression of genes is not the same throughout the body. Expression in blood and brain correlates only moderately (Pearson’s r of 0.24-0.64), with 35 to 80 % of genes being expressed in both tissues.1 Nevertheless, approximately 90 % of the weighted gene-gene interaction networks of the PFC transcriptome appear to be maintained in the peripheral blood. Brain and blood also show a significant overlap in terms of expressed quantitative trait loci, suggesting that common genetic effects (albeit of small effect size) may partially explain the comparability of gene expression in blood and brain. On this basis, computer models attempt to use blood gene expression to calculate gene expression in tissues.2
1.2. SNP: Single nucleotide polymorphisms
Single nucleotide polymorphisms (SNP, pronounced “snips”) are the most common form of genetic variation in humans. A SNP is a DNA deviation in one nucleotide. Example: at one point in the DNA, the nucleotide cytosine (C) is replaced by the nucleotide thymine (T).3
SNPs are found at around every 1,000th position in the entire human DNA. This means that there are around 4 to 5 million SNPs in the human genome.
To date, more than 600 million SNPs have been found worldwide.
For an SNP to be given a name (e.g. rs11420276), it must occur in at least 1 percent of the population.
Most SNPs are not found in the genes, but in the DNA between the genes.
Most SNPs have no effect on health or development. SNPs within a gene or in a regulatory region near a gene can influence the function of the gene and thereby play a more direct role in disease.
1.2. Alternative splicing (AS)
Alternative splicing (AS) is the process of splicing the precursor messenger RNA (pre-mRNA) and an important mechanism for increasing transcript diversity.4 AS is controlled by SNPs, which leads to variations in traits via complex mechanisms.5.
2. DNA (deoxyribonucleic acid)
A double-stranded concatenation of 4 possible nucleotides.
DNA is usually found in the cell nucleus as chromosomes (nuclear DNA = nDNA) and carries the genetic information of almost all living organisms. A small part of the DNA is located in the mitochondria (cellular power plants) as mitochondrial DNA (mtDNA).
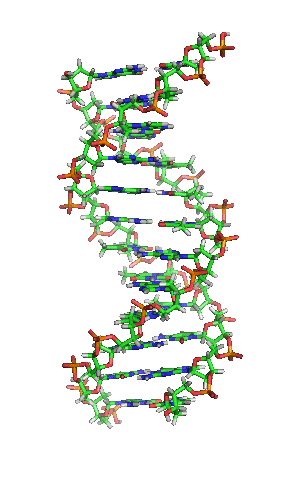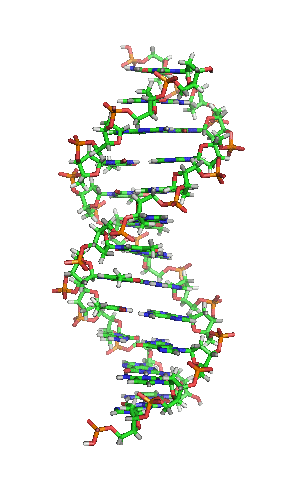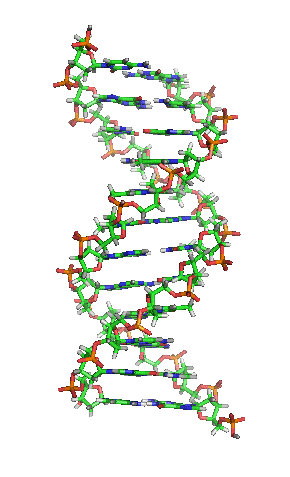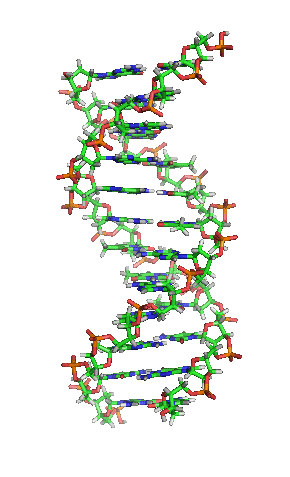
Model of a DNA. CC BY-SA 3.0
3. RNA (ribonucleic acid)
A mostly single-stranded concatenation of 5 possible nucleotide types.
RNA is the carrier of genetic information in certain viruses. There is RNA that passes on (genetic) information (coding RNA) and RNA that does not.
In 2017 and 2018 alone, 2000 new forms of RNA were discovered each year. PubMed found around 250,000 newly published scientific articles for the search term RNA in the years 2014 to 2018. By comparison, there were just under 11,500 for the keyword ADHD.
3.1. Coding RNA (mRNA)
mRNA copies the information contained in a gene and transmits it to the ribosome, where protein biosynthesis takes place using this information.
1.2 % of the RNA is large, protein-coding mRNA.
Forms:6
- NSD transcript
- Capped mRNA
- Consensus mRNA
- Edited mRNA
- Sample mRNA
- MRNA with frameshift
- Monocistronic mRNA
- Polyadenylated mRNA
- Polycistronic mRNA
- Recoded mRNA
- Trans spliced mRNA
3.2. Non-coding RNA (ncRNA)
There are many types of non-coding RNA.
ncRNAs are transcribed from DNA but not translated into proteins. They influence cell function and the pathogenesis of diseases by regulating gene expression through transcription and translation.7, e.g. through8
- Chromatinum construction
- Histone modification
- Binding of transcription factors
- MRNA degradation
- MRNA translation inhibition
- Interaction with splicing factors
- Interaction with RNA-binding proteins
As competitive endogenous RNAs (ceRNAs), they sequester miRNAs and modulate the expression of mRNAs.7
- Housekeeping ncRNA8
- are expressed in large quantities in all cells
- are involved in general cellular processes such as protein synthesis, RNA splicing and RNA modification
- ribosomal RNAs (rRNAs)
- Transfer RNAs (tRNAs)
- small nuclear RNAs (snRNAs)
- small nucleolar RNAs (snoRNAs)
- TERC (telomerase RNAs)
- tRNA-derived fragments (tRFs)
- tRNA halves (tiRNAs), while
- regulatory ncRNAs8
- regulate gene expression in a tissue-specific manner
- Micro-RNAs (miRNAs)
- small interfering RNAs (siRNAs)
- PI-WI-interacting RNAs (piRNAs)
- Enhancer RNAs (eRNAs)
- circular RNAs (circRNAs)
- long non-coding RNAs (lncRNAs)
- Y-RNAs (YRNAs)
- AsRNA (antisense RNA)
- Regulates gene expression
- Is transcribed from the coding DNA strand and not from the template DNA strand. It is therefore complementary to mRNA.
- CircRNA (circular RNA)7
- Binds to miRNA. Is thus involved in the regulation of cellular processes, such as proliferation or apoptosis (cell death)
- Regulation of gene expression, sponge miRNAs, modulation of neuronal development, synaptic plasticity 11
- Class I RNA
- Class II RNA
- EnhancerRNA
- Could have a regulatory function
- Guide RNA
- HnRNA (heterogeneous nuclear RNA)/ pre-mRNA (pre-mRNA, precursor mRNA)
- Precursor of the mature mRNA
- LncRNA (long non-coding RNA)
- Longer than 200 nucleotides
- Scaffold-DNA-chromatin complexes, inactivation of the X chromosome, telomere regulation, imprinting11
- long noncoding RNA7
- MiRNAs (microRNAs)
- Around 2,600 different miRNAs are known in humans8
- Closely related to siRNAs
- Regulate cellular processes such as proliferation and cell death with
- MiRNAs act as post-transcriptional gene silencers by interfering with target mRNAs and inhibiting their activity1213 MicroRNAs (miRNAs) thereby generally downregulate gene expression.
- The interaction between miRNAs and messenger RNAs (mRNAs) leads to the degradation or suppression of translation of the target mRNA. This makes them an essential factor in the post-transcriptional regulatory mechanism11
- In RNA interference (RNAi), the introduction of double-stranded RNA (dsRNA) blocks gene expression
- PiRNA (Piwi-interacting RNA)
- 26-31 nucleotides long
- Form complexes with PIWI proteins that are involved in epigenetic and post-transcriptional silencing in germ cells
- Transposon repression, DNA methylation11
- Riboswitches
- Serve to regulate genes
- Activating or repressing
- Serve to regulate genes
- Ribozymes
- Catalytically active RNA molecules
- Catalyze chemical reactions, just like enzymes
- RRNA (ribosomal RNA)
- Does not carry genetic information (like tRNA)
- Is involved in the assembly of the ribosome
- Is also catalytically active when the peptide bond is formed
- RRNA cleavage RNA
- RasiRNA (repeat associated small interfering RNA)
- RNase MRP RNA
- Essential for the catalytic activity of RNase MRP.
RNase MRP is an enzymatically active ribonucleoprotein that participates in the initiation of mitochondrial DNA replication in the mitochondria and is involved in precursor rRNA processing in the nucleus, where it splices the internal transcribed spacer 1 between 18S and 5.8S rRNAs.
- Essential for the catalytic activity of RNase MRP.
- RNase P RNA
- ScRNA (small cytoplasmic RNA)
- ScaRNA (small cajal body-specific RNA)
- Can be found in the Cajal Bodies
- Subclass of snoRNA
- Control the modification (methylation and pseudouridylation) of the RNA polymerase II-transcribed spliceosomal RNAs U1, U2, U4, U5 and U12.
- ScaRNA1 (small cajal body-specific RNA 1, ACA35)
- Appears to be involved in the pseudouridylation of U2-spliceosomal RNA at residue U89
- ShRNA
- SiRNA (small interfering RNA)
- Occurs in a cell signaling pathway known as RNAi (RNA interference)
In this process, dsRNA (double-stranded RNA) is divided into many small siRNAs by the enzyme Dicer and incorporated into RISC (RNA-induced silencing complex, an enzyme complex). Using the RNA fragments it contains, RISC binds to DNA (e.g. to gene regions) or to mRNA and can thus “switch it off”. - SiRNAs are involved in various cell processes and diseases.
- Occurs in a cell signaling pathway known as RNAi (RNA interference)
- small RNA7
- SnRNA (small nuclear RNA)
- Are found in the cell nucleus
- Responsible for splicing the hnRNA on the spliceosome.
- SnoRNA (small nucleolar RNA)
- Are found in the nucleolus
- Closely related to scaRNAs
- Small regulatory ncRNA
- SRP RNA
- SnRNA (small nuclear RNA)
- TRNA (transfer RNA)
- Does not encode genetic information
- Auxiliary molecule in protein biosynthesis
- Takes up individual amino acids from the cytoplasm and transports them to the ribosome
- TRNA is encoded by a specific RNA gene
- TasiRNA
- Telomerase RNA
- Telomeric transcript
- Three prime overlapping ncRNA
- TsRNA
- Regulation of gene expression, cell processes, stress reactions and the immune system11
- TSSa RNA
- Securing the transcription (?)11
- Vault RNA
- Y RNA
4. Nucleotides
Nucleotides each consist of a base, a sugar and a phosphate:
- Base (five possible bases)
- Adenine (A)
- Guanine (G)
- Cytosine (C)
- Thymine (T)
- Uracil (U) (in DNA only thymine instead)
- Sugar
- Ribose (D-ribofuranose) or
- Deoxyribose (2-deoxy-D-ribofuranose)
- Phosphate
- At least one phosphate group
5. Proteins (proteins)
Around 1 % of human genes encode proteins.
Proteins are necessary for the biological development of a living organism and for cell metabolism.
Human proteins can be formed from 21 different α-amino acids. The α-amino acids are linked together by peptide bonds to form a polymer (polypeptide). These polypeptides fold up into the native protein in an aqueous environment.
The biosynthesis of proteins takes place in all cells at the ribosomes using mRNA, which transmits the blueprint of proteins as genetic information.
The sequence of the bases of the mRNA encodes the sequence of the (proteinogenic) amino acid in triplets (codons). After translation, the side chains of some amino acids incorporated in the protein can still be modified.
One study found a correlation of ADHD with the proteins lysosomal Pro-X carboxypeptidase and alpha-2-antiplasmin in blood plasma.14
6. Amino acids (aminocarboxylic acids)
Amino acids are chemical compounds with an amino group (contains nitrogen) and a carboxylic acid group (contains carbon) that occur in all living organisms.
Amino acids are the building blocks of proteins.
They are primarily used to build up body tissue and are the final stages in the breakdown of protein.
Essential amino acids cannot be produced by an organism itself and must therefore be ingested through food.
Amino acids can be divided into different categories:
6.1. Types of amino acids
6.1.1. According to structure
- Amino acids by structure
- Α-Amino acids (2-aminocarboxylic acids, e.g. glycine)
- Β-amino acids (3-aminocarboxylic acids, e.g. β-alanine)
- Γ-Amino acids (4-aminocarboxylic acids, e.g. γ-aminobutyric acid = GABA)
6.1.2. Amino acids by function
- Proteinogenicity
Proteinogenicity means that the amino acid serves as a building block for proteins- Proteinogenic amino acids
- Α-amino acids, which are the building blocks of proteins, in humans 21 different
Name / abundance in proteins / essential:- Alanine (9.0 %)
- Arginine (4.7 %) partially essential
- Asparagine (4.4 %)
- Aspartic acid (5.5 %)
- Cysteine (2.8 %) essential for children and pregnant women
- Glutamine (3.9 %)
- Glutamic acid (6.2 %)
- Glycine (7.5 %)
- Histidine (2.1 %) partially essential
- Isoleucine (4.6 %) essential
- Leucine (7.5 %) essential
- Lysine (7.0 %) essential
- Methionine (1.7 %) essential
- Phenylalanine (3.5 %) essential
- Proline (4.6 %)
- Pyrrolysine (non-canonical, only in bacteria)
- Selenocysteine (non-canonical)
- Serine (7.1 %)
- Threonine (6.0 %) essential
- Tryptophan (1.1 %) essential
- Tyrosine (3.5%) essential for children and pregnant women
- Unknown amino acid (unclear whether amino acid)
- Valine (6.9 %) essential
- Α-amino acids, which are the building blocks of proteins, in humans 21 different
- Non-proteinogenic amino acids
- Naturally occurring (= biogenic / organogenic) amino acids
- More than 400 non-proteinogenic amino acids with biological functions
- D-amino acids (rare) are a special subgroup
- Synthetically produced / theoretically possible amino acids
- Considerably larger number
- Naturally occurring (= biogenic / organogenic) amino acids
- Proteinogenic amino acids
- Neurotransmitters
- Some amino acids act as neurotransmitters
- GABA
- Glycine
- As well as some degradation products of amino acids
- Some amino acids act as neurotransmitters
- Hormones
- Tissue mediators
6.1.3. Amino acids by origin
- Naturally occurring (= biogenic / organogenic)
- Synthetically produced
Tylee DS, Kawaguchi DM, Glatt SJ. On the outside, looking in: a review and evaluation of the comparability of blood and brain “-omes”. Am J Med Genet B Neuropsychiatr Genet. 2013 Oct;162B(7):595-603. doi: 10.1002/ajmg.b.32150. PMID: 24132893. REVIEW ↥
Hess JL, Quinn TP, Zhang C, Hearn GC, Chen S; Neuropsychiatric Consortium for Analysis and Sharing of Transcriptomes; Kong SW, Cairns M, Tsuang MT, Faraone SV, Glatt SJ. BrainGENIE: The Brain Gene Expression and Network Imputation Engine. Transl Psychiatry. 2023 Mar 22;13(1):98. doi: 10.1038/s41398-023-02390-w. PMID: 36949060; PMCID: PMC10033657. ↥
medlineplus.gov: What are single nucleotide polymorphisms (SNPs)? ↥
Wang J, Zhu QW, Mai JH, Zhang S, Wang Y, Liang J, Zhou JY (2024): A multi-omics study of brain tissue transcription and DNA methylation revealing the genetic pathogenesis of ADHD. Brief Bioinform. 2024 Sep 23;25(6):bbae502. doi: 10.1093/bib/bbae502. PMID: 39406522; PMCID: PMC11479714. ↥
Qi T, Wu Y, Fang H, Zhang F, Liu S, Zeng J, Yang J (2022): Genetic control of RNA splicing and its distinct role in complex trait variation. Nat Genet. 2022 Sep;54(9):1355-1363. doi: 10.1038/s41588-022-01154-4. PMID: 35982161; PMCID: PMC9470536. ↥
Gao Y, Takenaka K, Xu SM, Cheng Y, Janitz M (2025): Recent advances in investigation of circRNA/lncRNA-miRNA-mRNA networks through RNA sequencing data analysis. Brief Funct Genomics. 2025 Jan 15;24:elaf005. doi: 10.1093/bfgp/elaf005. PMID: 40251826; PMCID: PMC12008121. REVIEW ↥ ↥ ↥ ↥ ↥
Karaivazoglou K, Triantos C, Aggeletopoulou I (2025): Non-Coding RNAs in Neurodevelopmental Disorders-From Diagnostic Biomarkers to Therapeutic Targets: A Systematic Review. Biomedicines. 2025 Jul 24;13(8):1808. doi: 10.3390/biomedicines13081808. PMID: 40868063; PMCID: PMC12384018. REVIEW ↥ ↥ ↥ ↥
Khoodoruth MAS, Khoodoruth WNC, Uroos M, Al-Abdulla M, Khan YS, Mohammad F (2024): Diagnostic and mechanistic roles of MicroRNAs in neurodevelopmental & neurodegenerative disorders. Neurobiol Dis. 2024 Nov;202:106717. doi: 10.1016/j.nbd.2024.106717. PMID: 39461569. REVIEW ↥ ↥ ↥ ↥ ↥ ↥
Guo H, Ingolia NT, Weissman JS, Bartel DP (2010): Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature. 2010 Aug 12;466(7308):835-40. doi: 10.1038/nature09267. PMID: 20703300; PMCID: PMC2990499. ↥
Bartel DP (2009): MicroRNAs: target recognition and regulatory functions. Cell. 2009 Jan 23;136(2):215-33. doi: 10.1016/j.cell.2009.01.002. PMID: 19167326; PMCID: PMC3794896. REVIEW ↥
Cheng, Guan, Ma, Zhang, Cheng, Qi, Liang, Li, Kafle, Wen, Zhang (2020): An atlas of genetic correlations between psychiatric disorders and human blood plasma proteome. Eur Psychiatry. 2020 Feb 20;63(1):e17. doi: 10.1192/j.eurpsy.2019.6. PMID: 32093803; PMCID: PMC7315878. ↥